NIH’s All-of-Us Research Program Offers Free Genetic Testing to Increase Diversity of Its Database
All-of-Us program is free to participants and provides data to more than 800 research studies for cancer, COVID-19, Alzheimer’s, and other diseases; findings will lead to new biomarkers for clinical laboratory tests
It is hard to say no to free. At least that is what the National Institutes of Health (NIH) is counting on to help increase the size and diversity of its database of genetic sequences. The NIH’s All-of-Us Research Program is offering free genetic testing for all participants in the program, as well as free wearable Fitbits for those selected to provide lifestyle and behavior data.
Many pathologists and clinical laboratory managers know that this group of researchers hope to build a database of more than one million genetic sequences to better understand “how certain genetic traits affect underrepresented communities, which could greatly affect the future of customized healthcare,” CBS affiliate 8 News Now reported.
“Customized healthcare” is a euphemism for precision medicine, and genetic sequencing is increasingly playing a key role in the development of personalized diagnostics and therapeutics for cancer and other deadly diseases.
In “VA’s ‘Million Veterans Program’ Research Study Receives Its 100,000th Human Genome Sequence,” Dark Daily described how the NIH’s All-of-Us program was launched in 2018 to aid research into health outcomes influenced by genetics, environments, and lifestyle. At that time, the program had biological samples from more than 270,000 people with a goal of one million participants.
Matthew Thombs, Senior Project Manager of Digital Health Technology at Scripps Research in La Jolla, Calif., joined the All-of-Us program after losing a family member “to a condition I believe could have been managed with changes to their lifestyle,” he told 8 News Now.
“What we are building will empower researchers with the information needed to make such conclusions (about possible need to change lifestyles) and forever alter how diseases are treated,” he added. “I hope that what we are doing here will help my son grow up in a world where healthcare is more of a priority, and many of the ailments we see today are things of the past.”
Such genetic testing could discover biomarkers for future personalized clinical laboratory diagnostics and drug therapies, a key aspect of precision medicine.
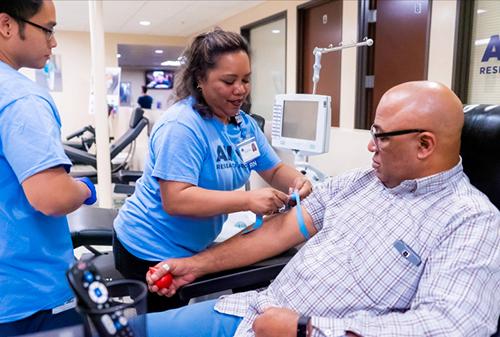
The photo above shows an All-of-Us participant being prepped to provide a biological sample for genetic testing. According to Matthew Thombs, Senior Project Manager of Digital Health Technology at Scripps Research, “participants can provide as much or as little information as they like, every single data point matters.” The collected data is shared anonymously with more than 800 research studies for COVID-19, Alzheimer’s, cancer, and other diseases, 8 News Now reported. (Photo copyright: KLAS-TV.)
Scripps Research Integrates Mobile Health Technology into All-of-Us Program
A critical aspect of the NIH’s research is determining how people’s behavior combined with their genetics may predispose them to certain diseases. Nonprofit research institution Scripps Research is working with the NIH’s All of Us Research Program to enroll and collect biological samples from one million US residents.
In addition, Scripps is fitting study participants with wearable mobile health devices to capture data on their habits and lifestyles.
“Until now, the treatment and prevention of disease has been based on a ‘one-size-fits-all’ approach, with most therapeutics tailored for the ‘average patient’. However, advances in genomic sequencing, mobile health technologies, and increasingly sophisticated informatics are ushering in a new era of precision medicine. This new approach takes into account differences in people’s genes, environment, and lifestyles giving medical professionals resources to design targeted treatments and prevention strategies for the individual,” Scripps states on its website.
Can wearable fitness devices and related data contribute to research on genetics and healthcare outcomes? Scripps aims to find out. It has fitted 10,000 people in the All-of-Us program with Fitbit devices (Fitbit Charge 4 tracker or Fitbit Versa 3 smartwatch) at no cost. Since February, Scripps has distributed 10,000 Fitbit wearable devices through the All-of-Us program.
“By sharing information about their health, habits, and environment, participants will help researchers understand why people get sick or stay healthy,” the Scripps website adds.
The Scripps researchers plan to analyze how the people use the wearable devices. They are also accumulating data about participants’ physical activity, heart rate, sleep, and other health metrics and outcomes “as part of the broader All of Us program,” a Scripps news release explained.
“This is the first time All of Us is distributing devices to participants. Our goal is to better understand how participants engage during research studies in order to continually improve user experience and participation. We also expect to learn more about how wearable data may inform the personalization of healthcare,” said Julia Moore Vogel, PhD, Director of The Participant Center at the All of Us Research Program at Scripps Research, in the news release.
All-of-Us Program Records ‘Significant Progress in Participant Diversity’
As of June, the NIH has enrolled 386,000 participants into the All-of-Us program, with 278,000 consenting to all of the program’s steps. Eighty percent of biological samples in the collection are from people in communities that have been under-represented in previous biomedical research an NIH new release noted. According to the NIH, that gives the All-of-Us research program “the most diverse dataset.”
What will all this research ultimately bring to clinical laboratories? Who knows? Nevertheless, if federal institutions like the NIH and non-profit research companies like Scripps believe precision medicine is worth investing in, then the All-of-Us program is worth watching.
A diverse database of a million genetic sequences combined with lifestyle and behavioral data may lead to new and improved personalized diagnostics and drug therapies.
—Donna Marie Pocius
Related Information
Free Genetic Testing Offered to Propel Medical Research; All of Us Building “Most Diverse Database”
NIH’s All of Us Research Program Records Significant Participant Diversity and Research Underway
VA’s Million Veterans Program Research Study Receives Its 100,000th Human Genome Sequence





