These new findings may affect how microbiology labs and physicians diagnose and treat several gastrointestinal conditions
Once again, a research effort has teased out new insights into the role the human microbiome plays in our digestive processes. Microbiologist and medical laboratory managers will be interested to learn that, according to the study team, specific microbes have a role in regulating how fast food moves through the digestive tract.
Researchers at the Dey Laboratory in Seattle recently examined the function of microbial bile acid metabolism in gut motility. They determined that “metabolites generated by the gut microbiome regulate gut transit,” according to a new paper published by the Fred Hutchinson Cancer Research Center (Fred Hutch).
“These findings have potential implications for the treatment of gastrointestinal conditions,” noted a Fred Hutch news release. This may mean new clinical laboratory tests to identify these strains of bacteria, along with new therapies for treating patients.
Gut motility (aka, Peristalsis) is the term used to describe the movement of food from the time it enters via the mouth until it leaves the body. This movement, the researchers found, is regulated by interactions between diet, the enteric nervous system (ENS) and the gut microbiota via processes that include bile acid metabolism.
Sex, Diet, and Lifestyle All Affect Treatment for Gastrointestinal Diseases
The Dey Laboratory researchers also discovered that sex was a significant variable in determining transit times with males having larger pro-motility effects.
In “Microbiome-encoded Bile Acid Metabolism Modulates Colonic Transit Times,” the Dey Laboratory researchers noted that previous studies have shown higher motility and varying bile acid profiles between men and women. They published their study in iScience, an open-access Cell Press journal.
“Our results suggest that strategies for treating or preventing gastrointestinal diseases may need to be tailored to sex and to biogeography of the gut,” they wrote. “While targeting the microbiome and the ENS is justified, our observation of significant transcriptional responses to defined interventions in a highly controlled gnotobiotic setting also highlights challenges to clinical translation.”
The researchers concluded that:
- Gut microbiome-generated bile acids regulate colonic transit via TGR5 protein.
- Lithocholic acid (LCA) had the largest colonic pro-motility effect.
- Bile acids exert sex-biased effects on gut transit times.
- Enteric nervous system (ENS) transcriptional responses are regional- and microbiome-specific.
“The human experience—which reflects the aggregate effects of the innumerable dietary ingredients that we consume daily, the hugely diverse metabolically dynamic microbes that inhabit our guts, our own digestive processes, and the interactions of all of the above that result in thousands of gut metabolites—entails significantly more complex and variable transcriptional responses to environmental cues,” the Dey Laboratory scientists concluded.
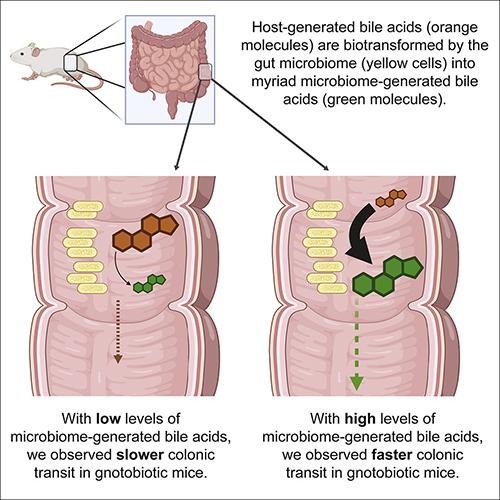
To perform their research, the scientists developed both high and low BSH (bile salt hydrolase) bacterial communities for germ-free mice, which are known to exhibit slower gut motility and less complex bile acid profiles than colonized animals. (See graphic above taken from the Dey Laboratory published paper.)
The spice turmeric and dyes were added to the diets of the mice to track gut motility. The mice that were given the BSH-high microbiota had higher fecal concentrations of unconjugated bile acids than those given the BSH-low form of the microbiota. The mice given the BSH-high version also experienced faster transit times, according to the researchers’ iScience paper.
The researchers also concluded that the BSH-high group had greater fecal concentrations of lithocholic acid (LCA) which indicates variations in bile acid metabolism might affect gut transit.
When the scientists infused bile acids directly into mouse colons, variable acids reacted differently with LCA having the fastest transit times. The researchers hypothesized that LCA might signal through a bile receptor known as TGR5 which blocked the effects of LCA on colonic transit times. TGR5, also called G protein-coupled bile acid receptor, functions as a cell surface receptor for bile acids.
The Dey Laboratory team developed a method to measure expression changes in ENS genes and found that neither BSH activity nor gut transit phenotypes were major drivers of gene expression changes. They found that the location of the gut segment, or biogeography, was the leading contributor to ENS signature variance between samples.

“We expected to see shared host transcriptional responses in mice harboring communities with similar metabolic profiles. However, we did not see this for the most part,” explained gastroenterologist Neelendu Dey, MD (above), a physician/scientist and Assistant Professor, Clinical Research Division, at Fred Hutchinson Cancer Research Center, in the press release. “If anything, shared responses were regional, and these signatures did not cluster by BSH/motility phenotypes.” (Photo copyright: Seattle Cancer Care Alliance.)
The scientists “identified consortium-specific transcriptional changes in genes involved in ENS signaling, development, maintenance, and bile acid metabolism, and these differed across regions of the GI tract. Together these findings indicate that ENS transcriptional responses are regional and microbiome-specific,” according to the Fred Hutch press release.
“This remains a confusing part of the story for us—how is it that we can see predictable host motility responses when colonizing the guts of gnotobiotic mice with phenotypically defined communities, but the middle-man (the host enteric nervous system) appears to have such varied responses?” the Dey Laboratory researchers noted in the press release.
“It suggests that gut motility phenotypes that appear similar may in fact represent (when we look under the hood) diverse host physiologic phenotypes that we are just beginning to understand,” they added.
The results of this study could have potential implications for the precision medicine diagnosis and treatment of gastrointestinal illnesses.
Blue Poop Challenge
Earlier this year, people were encouraged to participate in the “blue poop challenge” conducted by research company ZOE Global Limited (ZOE) to determine how long it takes food to travel through the body.
ZOE is also known for collaborating with King’s College London, and Guy’s and St Thomas’ Hospitals to create the COVID Symptom Tracker mobile app (now known as the COVID Symptom Study).
For the Blue Poop Challenge, individuals are asked to eat blue muffins and then report on the company’s website as to how long it took for the blue dye to appear in their stools.
The purpose of this ongoing study is to reveal pertinent information about an individual’s gut health and microbiome.
Since 2010, Dark Daily has reported on dozens of research studies and innovative developments involving human microbiome and gut bacteria and their critical importance in the development of clinical laboratory testing, drug therapies, and precision medicine.
In “University of Utah and Sloan Kettering Institute Study Sheds Light on How the Body Recognizes ‘Good’ from Bad Bacteria in the Microbiome,” we reported on research being conducted at the University of Utah and the Sloan Kettering Institute (SKI) which found that early in life intestinal microorganisms “educate” the thymus to develop T cells.
These studies’ findings could lead to improved immune system therapeutics and associated clinical laboratory tests.
“All of this suggests the potential in the future for clinical laboratories and microbiologists to do microbiome testing in support of clinical care,” said Robert Michel, Editor-in-Chief of Dark Daily and its sister publication The Dark Report.
More research is needed in these areas. But gut bacteria and the human microbiome are an integral part of our health and wellbeing. It is worth keeping an eye on new developments in those fields of study.
—JP Schlingman
Related Information
Keeping Regular: Gut Bacteria Modulate Transit Time via Bile Acids
Microbiome-encoded Bile Acid Metabolism Modulates Colonic Transit Times
Does the Viral Blue Poop Challenge Really Tell You Anything about Gut Health?
The Blue Poop Challenge Could Tell You Important Info about Your Gut Health—Here’s How It Works